
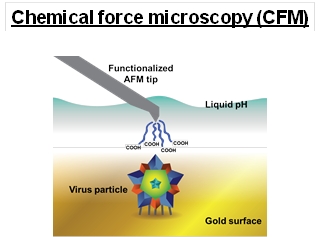
Virus surface chemistry plays a key role in virus sorption and partitioning processes, however there is very limited information on viral surface chemical properties. To enhance the toolbox of available characterization tools, chemical force microscopy (CFM) is being pioneered for viral particle characterization. CFM measures the adhesion force between a particle, in this case a virus particle, and a functionalized atomic force microscopy (AFM) tip. CFM reduces many of the difficulties of bulk characterization techniques by measuring adhesion of individual virus particles, likely reducing purification affects that can change surface properties. Two important surface characteristic that can dominate separation processes are isoelectric point (pI) and hydrophobicity. The surface charge of virus particles is pH dependent, which dictates the mobility and controls the colloidal behavior in virus sorption processes. While traditional characterization methods, including zeta potential and viral adsorption to a charged surface chemistry, have been used to study viral surface charges and determine the pI of virus, they are bulk measurements that can change when the studied particle has a different purification method. Hydrophobic interactions are known to be strong for virus particles and are implicated to be important in virus entry into cells. The single particle method of CFM allows not only the quantitation of the virus surface chemistry, but could be used to understand and better design purification processes. The adhesion of a viral particle to any chemical moiety in any solution condition can be tested with this method. The model non-enveloped porcine parvovirus (PPV) and enveloped bovine viral diarrhea virus (BVDV) were used to demonstrate the use of CFM to determine the hydrophobicity and pI for viral particles. This information is being used to determine the binding and elution conditions of a virus to a chromatographic column. With a thorough understanding of virus surface characteristics, virus sorption and desorption processes could be significantly improved and designed to specifically target viral particles using minimal viral material during development.
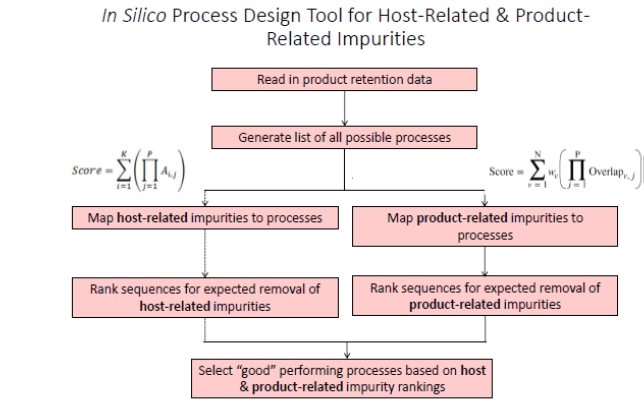
Designing integrated non-affinity downstream processes for biologics poses a significant challenge due to the broad range of design space available for resin selection and buffer conditions. To address this challenge, we developed an in silico-based approach to quickly design and rank a fully inclusive list of integrated downstream processes for their ability to remove impurities using only orthogonally selective multimodal and ion exchange resins. This approach involves the one-time characterization of an impurity database and considers both impurity profiles patterns and product retention behavior to generate and score all possible integrated purification trains. This database was then employed in concert with an in silico process development tool which generates and ranks all possible integrated chromatographic sequences for their ability to remove orthogonal impurities. Top-ranking outputs are then used to guide the experimental development and refinement of purification processes, significantly expediting the development of downstream processes. For Pichia pastoris, this approach was used for two non-mAb products, hGH and G-CSF, which were successfully purified from cell culture fluid process impurities in a process that resulted in high purity and product recovery. We also performed modifications to our process design approach in order to account for both process-related impurities and product-related impurities. This modified strategy was first shown to be successful for purifying IFN produced in Pichia pastoris, and was then applied to the non-affinity-based purification of a mAb-aggregate challenge in CHO, an expression system with a much higher HCP burden. Finally, results are presented on a mathematical framework to characterize and quantify orthogonality in multimodal systems. Using this, we observed several interesting and unexpected results including the existence of a highly orthogonal pair of resins belonging to the same class/family.
Bivalent bispecific antibodies (BiS) are produced by recombinantly expressing a whole monoclonal antibody (mAb) with two single chain variable fragments (scFv) using flexible polypeptide linkers. Three homologous BiS samples varying only by scFv attachment location show reversible, three-peak elution profiles on ProPac WCX-10 and Source 15S cation exchange columns. Collection and reinjection of any of the three peaks results in the same three-peak elution profile. This behavior is seen at pH values in the range 5–8.5. In-line DLS showed that all peaks have a monomer-sized hydrodynamic radius of 6.2 nm. Unlike the normal chromatographic behavior, decreasing flow rate decreased resolution. At the lowest flow rate tested, only a single symmetric peak was observed eluting between the first and second peak of the higher flow rate elution profiles. A hold step at strong binding conditions followed by gradient elution with high flow rate enriched the third, most retained, peak. Biophysical measurements including in-line fluorescence, chemical cross-linking, enzyme digestion to determine domain contributions, and a mechanistic model reveals that the three-peak elution is caused by conformational interconversion occurring on time sales comparable with chromatographic separation. From the peptide sequence, the scFvs carry a net positive charge. When the scFv is folded onto the base mAb weaker bind to the resin occurs whereas if the scFv is outstretched more favorable interaction and stronger binding occurs on the resin surface. The three peaks from weakest to strongest retention are: both scFvs interacting with base mAb, one scFv interacting with base mAb and one outstretched, and both scFvs outstretched. Equilibrium favors the tight conformation in solution and outstretched while adsorbed. The highly flexible linkers allow slow changes in tertiary structure responsible for reversible, three-peak elution behavior. Analogous results are observed on HIC columns suggesting that similar molecular interactions are responsible for the multi-peak elution behavior.
Recently, exosomal vesicles are known as a next generation of biomarkers for molecular diagnosis, and their separation methods has been studied last decades. T-cell Immunoglobulin and Mucin domain (TIM) which is Ca-dependent membrane protein has an ability to interact phosphatidyl serine (PS) clusters on inner cell membrane, and thus it is considered that TIM fragment is one of ligand candidates to separate exosomal vesicles from serum as well as culture supernatant. Also, Exosome can be released from the resin.
Here, we designed and characterized TIM fragments genetically-fused with Avi-tag and PMMA-tag for site-direct immobilization to solid supports.
Avi-tag-fused TIM fragments were successfully produced by HEK293T cells with co-expression with Biotin ligase BirA. Consequently, site-specific biotinylation at the C-terminal region of TIM fragments could be achieved. Mono-biotinylated TIM fragment (ABT-TIM-VM) could be immobilized on the surface of streptavidin-coated plate, and could recover 10-times lower concentration of exosomes, compared with the plate with randomly-biotinylated TIM (RBT-TIM-VM). These results indicated that orientation control of TIM on the surface of solid support is more important to improve recovery of exosomes. Furthermore, biotinylation yields of ABT-TIM-VM could be controllable by changing the ratio of expression vectors for TIM and BirA.
Another candidate, PMMA-tag was introduced at the C-terminal region of TIM fragments. TIM-VM-PM could also be produced by 293T cells, and purified by Ni-NTA chromatography. The optimal buffer condition for TIM-VM-PM was screened from 96 conditions with different pH and NaCl concentrations. Consequently, 25-times higher signal intensity of sandwich ELISA for detection of exosome was detected, compared with the plate with tagless TIM-VM. These results indicated that PMMA-tag introduced was functional and preferentially adsorbed the surface of PMMA plate.
Thus, both TIM fragments developed in this study, namely, ABT-TIM-VM and TIM-VM-PM are useful ligand proteins to separate exosomal vesicles from biological samples.
The synthetic DNA containing modified nucleotides comprise a powerful toolbox of manipulating the formation of nucleic acid duplexes for diagnosis and therapeutic purposes. We are interested in the DNA oligonucleotides containing site-specific neutral internucleoside methyl phosphotriester (MPTE) linkages, nDNA in short (n for neutral). nDNA is interesting because the electrostatic repulsion between nDNA and its complementary strand would decrease and the duplex formation could be enhanced. However, the methyl group of MPTEs may sterically hinder duplex formation and destabilize the duplexes. Therefore, the uses of nDNA may offer additional elements of maneuvering the stability of nucleic acid duplexes formation. The structural information and hybridization thermodynamics were reported in this study. Based on the structural and thermodynamics information of nDNA with natural DNA, successful results in the molecular detection method such as PCR and qPCR will be reported. The applications of nDNA on the in-situ hybridization will be discussed as well. Most importantly, using nDNA on the semi-conductor based biochip and biosensor, such as field effect transistor, has generated wide avenue for the precision medicine, which also will be highlighted in this report.
Keywords: methyl-phosphotriester linkage of DNA (nDNA); molecular detections; PCR and qPCR; In-situ hybridization; field effect transistor